Par Benjamin Louis | 3 Apr 2019 |

Quels outils pour quel résultat
En plus de packages spécifiques présentés plus tard, ce travail est avant tout un travail de manipulation de données et évidemment les packages du tidyverse vont nous être utiles : dplyr pour la manipulation de données en général, purrr pour la programmation fonctionnelle et stringr pour la manipulation de chaîne de caractères.
library(dplyr)
library(purrr)
library(stringr)Le questionnaire Google original étant confidentiel, j’en ai créé un factice en mixant plusieurs modèles proposés par Google. J’ai pu facilement télécharger le fichier PDF correspondant. Pour la version HTML, j’ai d’abord essayer d’atteindre directement la page depuis la version éditable du questionnaire à l’aide du package rvest, mais j’ai perdu la bataille contre la double authentification de Google. Si quelqu’un a une solution pour ce problème, je suis preneur ! Tant pis pour le classique webscraping, j’ai opté pour la sauvegarde de la page HTML depuis mon navigateur pour travailler en local. L’ensemble des documents est disponible ici.
À partir de ces fichiers, je cherche à obtenir un data.frame final contenant 3 colonnes :
questions: les titres des questionstypes: le type des questions qui peut prendre 4 valeurs :uniquepour des questions à choix multiple avec une seule réponse possible,multiplepour des questions à choix multiple avec plusieurs réponses possibles,freepour les questions avec réponse libre etgridpour les grilles à choix multiples.choices: une liste-colonne dont chaque ligne est un vecteur des choix possibles de réponse pour les questionsuniqueetmultiple, un vecteur dont les éléments sont nommés pour les questionsgrid(les noms correspondant à ligne1, ligne2, …, colonne1, … de la grille) et un vecteur de longueur 0 pour les questionsfree.
pdftools à l’essai
Pour travailler avec le fichier PDF, j’utilise le package rOpenSci pdftools développé par Jeroen Ooms’. Si vous ne connaissez pas ce package, je vous encourage vivement à y jeter un oeil ainsi qu’à l’article de Maëlle Salmon sur l’emploi du temps de Trump. Pour les utilisateurs d’Ubuntu, l’installation de pdftools peut s’avérer délicate car la dernière version de Poppler n’est pas disponible sur Linux. J’ai réussi grâce à cette discussion (testé avec Ubuntu 18.04).
Importer les données d’un PDF
La fonction pdf_data du package pdftools est particulièrement utile pour importer des donnnées depuis un PDF. Cette fonction renvoie une liste de data.frame, un par page du document, contenant une ligne par “boîte” contenant du texte (i.e. texte entre deux zones sans texte tel que des espaces) et les colonnes correspondant à la taille, aux coordonnées et à la valeur de ces “boîtes”.
library(pdftools)
metapdf <- pdf_data("2019-04-03-pdf-vs-html_files/test_form.pdf")
head(metapdf[[1]])## # A tibble: 6 x 6
## width height x y space text
## <int> <int> <int> <int> <lgl> <chr>
## 1 49 11 0 0 FALSE Introduction
## 2 4 11 0 781 TRUE 1
## 3 12 11 7 781 TRUE sur
## 4 4 11 22 781 FALSE 2
## 5 287 11 323 0 FALSE https://docs.google.com/forms/d/10UqbE6edL7K6G~
## 6 100 19 77 80 FALSE IntroductionLa fonction est vraiment parfaite pour ce travail car le format de l’objet qu’elle renvoie est très proche du format final désiré. Il suffit de construire un unique data.frame à partir de tous ceux présents dans la liste en prenant soin de garder un identifiant des pages du document.
library(pdftools)
metapdf <- bind_rows(metapdf, .id = "page")Et voilà ! Avec cette unique fonction utilisée une seule fois, toutes les données désirées ont été extraites du PDF. Il ne reste plus qu’à remanier les données… ce que j’imaginais très simple avant de vraiment me plonger dedans ! Nettoyer ces données signifient ici trouver des règles pour éliminer ce qui ne nous intéressent pas et mettre en bon format ce qui nous intéressent. En pratique, trouver ces règles n’a pas été aussi simple que j’imaginais.
Nettoyage
Rassembler les lignes
Il faut d’abord sélectionner toutes les boîtes de texte dont la hauteur est égale à 10, hauteur qui semble être la hauteur pour les intitulés des questions et les réponses possibles. Le tableau de données qui a une ligne par boîte de texte est ensuite réduit à une ligne par ligne dans le PDF. Pour cela, l’hypothèse qui semble raisonnable est que les textes qui appartiennent à la même ligne du document ont une coordonnée y égale sur une même page. Les textes partageant la même coordonnée y sont donc réunis dans une nouvelle colonne nommée lines.
metapdf <- metapdf %>%
filter(height == 10) %>%
group_by(page, y) %>%
mutate(lines = str_c(text, collapse = " ")) %>%
distinct(lines, .keep_all = TRUE) %>%
ungroup()Identifier les phrases
La deuxième étape consiste à identifier les “phrases” présentent dans les lignes. L’idée générale ici est qu’une phrase commence soit par un ou deux chiffres suivis par un point puis un espace (pour les intitulés des questions) soit par une lettre majuscule suivie d’une lettre minuscule ou d’un espace (pour les réponses possibles). Il faut également pouvoir identifier les phrases qui se retrouvent sur plusieurs lignes. La colonne sentences_group est crée en a) détectant chaque boîte de texte commençant une phrase à l’aide d’expressions régulières, b) en appliquant une somme cumulée sur le booléen ainsi obtenu afin que chaque ligne appartenant à la même phrase ait la même somme et c) en ajoutant le prefix sent à cette somme. Cette colonne permet de réunir les phrases présentes sur plusieurs lignes et de supprimer celles qui sont présentent plusieurs fois.
metapdf <- metapdf %>%
mutate(sentences_group = str_detect(lines,"(^\\d{1,2}\\.\\s|^[:upper:]([:lower:]|\\s))") %>%
cumsum() %>%
paste0("sent", .)) %>%
group_by(sentences_group) %>%
mutate(sentences = str_c(lines, collapse = " ")) %>%
distinct(sentences_group, .keep_all = TRUE) %>%
ungroup()
select(metapdf, lines, sentences_group, sentences)
## # A tibble: 44 x 3
## lines sentences_group sentences
## <chr> <chr> <chr>
## 1 Event Timing: January 4th-~ sent1 Event Timing: January 4th-6th, 2~
## 2 Event Address: 123 Your St~ sent2 Event Address: 123 Your Street Y~
## 3 Contact us at (123) 456-78~ sent3 Contact us at (123) 456-7890 or ~
## 4 1. Name * sent4 1. Name *
## 5 2. Email * sent5 2. Email *
## 6 3. Organization * sent6 3. Organization *
## 7 4. What days will you atte~ sent7 4. What days will you attend? *
## 8 Check all that apply. sent8 Check all that apply.
## 9 Day 1 sent9 Day 1
## 10 Day 2 sent10 Day 2
## # ... with 34 more rowsIdentifier les blocs de question
La méthode précédente aboutissant à la colonne sentences_group est également utilisée pour créer la colonne question_bloc qui identifie les lignes qui appartiennent à la même question i.e. l’intitulé et les réponses possibles. Les lignes ayant la valeur bloc0 correspondent au texte d’introduction avant les questions et sont donc supprimées.
metapdf <- metapdf %>%
mutate(question_bloc = paste0("bloc", cumsum(str_detect(lines, "^\\d{1,2}\\.\\s")))) %>%
filter(question_bloc != "bloc0")Obtenir le type de question
Pour chaque bloc de question, le type de question associé est obtenu grâce à un comportement par défaut des formulaires Google qui ajoute une phrase après chaque intitulé de question pour préciser combien de réponses sont possibles.
metapdf <- metapdf %>%
group_by(question_bloc) %>%
mutate(types = case_when("Check all that apply." %in% sentences ~ "multiple",
"Mark only one oval." %in% sentences ~ "unique",
"Mark only one oval per row." %in% sentences ~ "grid",
length(sentences) == 1 ~ "free")) %>%
filter(!is.element(sentences, c("Check all that apply.", "Mark only one oval.",
"Mark only one oval per row."))) %>%
mutate(id = 1:n()) Isoler les questions et réponses possibles
L’identifiant précedemment créé (id) permet la différenciation entre les intitulés des questions (id == 1) et les réponses possibles (id != 1). Les questions de type grid ont besoin d’un traitement particulier, id == 2 correspondant aux noms des colonnes tandis que les autres id correspondent aux noms des lignes. Ces valeurs sont d’abord isolées puis les colonnes choices and questions sont créées pour finalement réunir de nouveau les réponses possibles des questions de type grid.
# Grid question
columns <- metapdf %>%
filter(types == "grid" & id == 2) %>%
pull(sentences) %>%
str_split("\\s") %>%
unlist()
rows <- metapdf %>%
filter(types == "grid" & !id %in% 1:2) %>%
pull(sentences)
# Creating questions and choices columns
metapdf <- metapdf %>%
mutate(choices = list(sentences[-1])) %>%
ungroup %>%
filter(id == 1) %>%
select(questions = sentences, types, choices)
# Adding choices values for grid question
metapdf$choices[metapdf$types == "grid"] <- list(c(row = rows, column = columns))Nettoyage
La dernière étape correspond à du nettoyage. Le symbole * est retiré des intitulés de question où la réponse est obligatoire.
metapdf <- metapdf %>% mutate(questions = str_replace(questions, "\\*", "") %>% str_trim)
metapdf
## # A tibble: 9 x 3
## questions types choices
## <chr> <chr> <list>
## 1 1. Name free <chr [0]>
## 2 2. Email free <chr [0]>
## 3 3. Organization free <chr [0]>
## 4 4. What days will you attend? multiple <chr [3]>
## 5 5. Dietary restrictions unique <chr [6]>
## 6 6. I understand that I will have to pay $$ upon arrival multiple <chr [1]>
## 7 7. What will you be bringing? multiple <chr [5]>
## 8 8. How did you hear about this event? unique <chr [5]>
## 9 9. What times are you available? grid <chr [9]>rvest à l’essai
Si vous n’avez jamais entendu parler de HTML et CSS, il est possible que ce qui va suivre sera un peu obscure. Le contenu d’une page web est stocké en HTML et il est possible de récupérer ce contenu à l’aide de package tel que rvest (c’est ce qu’on appelle du webscrapping).
rvest, selectorGadget et les sélecteurs CSS
De manière très brève, une page HTML a une structure en arbre (appelé un DOM pour Document Object Model) où chaque noeud contient des objets. La fonction read_html permet de lire et importer le DOM dans R.
library(rvest)
form <- read_html("2019-04-03-pdf-vs-html_files/test_form.html")
form
form %>% html_nodes("body") %>% html_children() %>% length()## {html_document}
## <html lang="en" class="freebird">
## [1] <head>\n<meta http-equiv="Content-Type" content="text/html; charset=UTF-8 ...
## [2] <body dir="ltr" class="freebirdLightBackground isConfigReady" id="wizView ...
## [1] 23Le premier noeud de notre page est appelé html et contient deux noeuds enfants appelés respectivement head et body. Si on jette un oeil aux noeuds enfants de body, on voit qu’il en existe 23. Ces noeuds peuvent également avoir des noeuds enfants et ainsi de suite.
La structure d’un noeud ressemble à <name attr1 = "value1" attr2 = "value2"> object </name> où <name>...</name> est une balise (avec un nom spécifique) qui peut avoir plusieurs attributs (attr1, attr2) ayant des valeurs spécifiques ("value1, "value2"). Les balises délimitent des objets qui peuvent être d’autre noeuds ou du texte présent sur la page HTML.
Les attributs des noeuds peuvent être des sélecteurs CSS. Ils sont utilisés pour lier les noeuds aux feuilles de style CSS qui définissent comment le contenu d’un noeud est affiché sur le navigateur web. Le nom de ces attributs est soit id ou class. Les sélecteurs CSS peuvent être utilisés comme des identifiants des noeuds lorsque l’on récupère du contenu web.
Le principal défi est d’utiliser le bon sélecteur CSS. Pour cela, l’outil SelectorGadget est très utile. Comme l’écrit Hadley Wickham dans cette vignette du package rvest, “Selectorgadget is a javascript bookmarklet that allows you to interactively figure out what css selector you need to extract desired components from a page.” (SelectorGadget est un outil javascript qui permet de distinguer interactivement les sélecteurs CSS nécessaires pour l’extraction d’un contenu désiré dans une page). SelectorGadget est facile d’utilisation et la vignette l’explique suffisamment clairement pour que je ne le détaille pas ici.
Récupérer les données
blocs de question
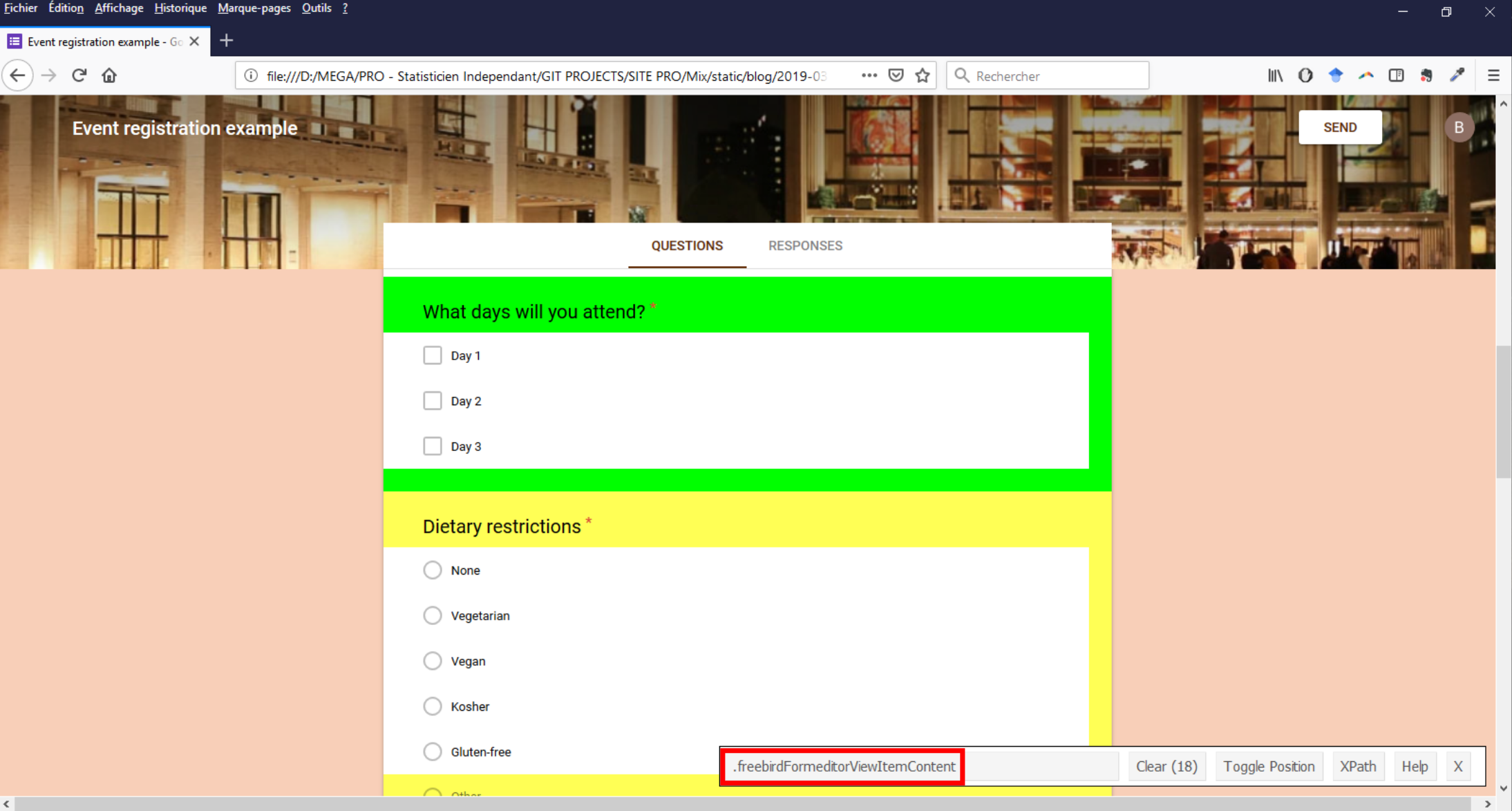
Les blocs de question (intitulé + réponses possibles) sont contenus dans des noeuds ayant comme classe CSS freebirdFormeditorViewItemContent.
Dans cette image (cliquer pour agrandir), GadgetSelector nous dit qu’il y a 18 noeuds ayant cette classe alors qu’il y seulement 9 questions dans le questionnaire. Je n’ai pu précisément identifier à quoi correspondent les 9 supplémentaires mais il s’avère que les 9 qui nous intéressent contiennent tous un noeud avec une classe CSS égale à freebirdFormeditorViewItemMinimizedTitleRow qui lui-même contient un texte correspondant à l’intitulé de la question.
prefix <- function(.x) {paste0(".freebirdFormeditorView", .x)}
question_blocs <- form %>%
html_nodes(prefix("ItemContent")) %>%
keep(~html_nodes(.x, prefix("ItemMinimizedTitleRow")) %>%
length() != 0)Intitulé des questions
Comme précédemment énoncé, l’intitulé des questions sont stockés dans des noeuds avec la classe freebirdFormeditorViewItemMinimizedTitleRow.
questions <- question_blocs %>%
html_nodes(prefix("ItemMinimizedTitleRow")) %>%
html_text()
questions
## [1] "Name*"
## [2] "Email*"
## [3] "Organization*"
## [4] "What days will you attend?*"
## [5] "Dietary restrictions*"
## [6] "I understand that I will have to pay $$ upon arrival*"
## [7] "What will you be bringing?*"
## [8] "How did you hear about this event?*"
## [9] "What times are you available?*"Réponses possibles et type de question
La prochaine étape consiste à isoler les réponses possibles et les types des questions en appliquant un fonction sur chaque bloc de question. Cette fonction contient des conditions de type if..else... qui dépendent du type de la question.
Les deux types de question à choix multiple sont dans des noeuds avec la classe freebirdFormeditorViewQuestionBodyChoicelistbodyOmniList. Les réponses possibles sont stockées dans les attributs value des noeuds ayant pour classe exportInput et l’attribut aria-label égal à "option value". Certaines de ces questions permettent de répondre une option autre (“Other”) - cette option nécessite un traitement spécial car stockée dans un autre type de noeud. De plus, le choix des types de question entre unique et multiple dépend de la valeur de l’attribut data-list-type des noeuds ayant comme classe freebirdFormeditorViewQuestionBodyChoicelistbodyOmniList.
Les questions de type Grid sont dans des blocs de question qui contiennent un noeud avec la classe freebirdFormeditorViewQuestionBodyGridbodyRow. Le texte des noeuds ayant uniquement cette classe correspond aux noms des lignes de la grille. Le nom des colonnes est stocké comme texte dans un noeud avec la classe freebirdFormeditorViewQuestionBodyGridbodyCell présent dans des noeuds avec la classe freebirdFormeditorViewQuestionBodyGridbodyColumnHeader.
# To make code smaller
get_xpath <- function(.y) {
cl <- str_replace(prefix(.y), "\\.", "")
paste0('.//*[@class="', cl,'"]')
}
# To get choices and types
get_choices_and_types <- function(.nd) {
# Multiple choices questions
if (length(html_nodes(.nd, prefix("QuestionBodyChoicelistbodyOmniList"))) != 0) {
# Answer choices
choices <- html_nodes(.nd, ".exportInput") %>%
keep(~html_attr(.x, "aria-label") == "option value") %>%
html_attr(.,"value")
# For "Other" option
other <- html_nodes(.nd, prefix("OmnilistListPlaceholderOption")) %>%
html_attr(., "style") != "display:none"
if (other) choices <- c(choices, "Other:")
# Question type
dltype <- html_nodes(.nd, prefix("QuestionBodyChoicelistbodyOmniList")) %>%
html_attr("data-list-type")
if (dltype == "1") type <- "multiple" else type <- "unique"
# Grid question
} else if (length(html_nodes(.nd, prefix("QuestionBodyGridbodyRow"))) != 0) {
# Answer choices
choices <- c(
row = html_nodes(.nd, xpath = get_xpath("QuestionBodyGridbodyRow")) %>%
html_text(),
column = html_nodes(.nd, prefix("QuestionBodyGridbodyColumnHeader")) %>%
html_nodes(xpath = get_xpath("QuestionBodyGridbodyCell")) %>%
html_text()
)
# Question type
type <- "grid"
# Free question
} else {
choices <- character(0)
type <- "free"
}
return(tibble(types = type, choices = list(choices)))
}
choices_types <- map_dfr(question_blocs, get_choices_and_types)Nettoyage
Finalement, la dernière étape correspond à relier les différentes données et d’appliquer le même nettoyage que pour le fichier PDF.
metahtml <- bind_cols(questions = paste0(1:9, ". ", unlist(questions)), choices_types)
metahtml <- mutate(metahtml, questions = str_replace(questions, "\\*", "") %>% str_trim)
metahtml
## # A tibble: 9 x 3
## questions types choices
## <chr> <chr> <list>
## 1 1. Name free <chr [0]>
## 2 2. Email free <chr [0]>
## 3 3. Organization free <chr [0]>
## 4 4. What days will you attend? multiple <chr [3]>
## 5 5. Dietary restrictions unique <chr [6]>
## 6 6. I understand that I will have to pay $$ upon arrival multiple <chr [1]>
## 7 7. What will you be bringing? multiple <chr [5]>
## 8 8. How did you hear about this event? unique <chr [5]>
## 9 9. What times are you available? grid <chr [9]>Conclusion
On peut commencer par vérifier que les deux tableaux de données metapdf et metahtml correspondent.
map2(metapdf, metahtml, all.equal)
## $questions
## [1] TRUE
##
## $types
## [1] TRUE
##
## $choices
## [1] TRUEAvec deux types différents de fichiers, il a été possible d’obtenir le même résultat grâce à deux packages vraiement utiles : pdftools et rvest. Récupérer les données depuis la page HTML aurait pu être effectuée avec le package xml2 car beaucoup des fonctions de rvest utilisées reposent sur les fonctions de xml2. La principale différence est que xml2 accepte uniquement des expressions xpath qui de mon point de vue ont une syntaxe plus difficile à maitriser. Cependant, il est possible d’obtenir les expressions xpath depuis GadgetSelector.
L’objectif de ce travail n’était pas de définir si un des deux moyens d’extraire des données est meilleur que l’autre. J’ai pris du plaisir dans les deux cas et j’ai été plutôt surpris par la puissance des deux packages utilisés. Lorsque que j’aurais besoin de travailler à partir de fichiers PDF, je pourrais compter sur pdftools sans problème, autant que je pourrais compter sur rvest pour extraire des données depuis le web.
S’il fallait pointer une différence, c’est que les règles utilisées pour arranger les données extraites du PDF dépendent fortement de la structure du PDF. La grande diversité des structures possibles rend probablement difficile la définition d’une stratégie reproductible tandis que pour les pages HTML, la stratégie est toujours la même : trouver les valeurs des sélecteurs CSS pour les données voulues à l’aide de SelectorGadget et appliquer la fonction appropriée pour les extraire. Au final, ce n’est peut être pas plus rapide mais cela rend le flux de travail plus évident.
J’espère que vous avez pris plaisir à lire ce premier article autant que j’en ai pris à l’écrire. N’hésitez pas à laisser un commentaire et dites-moi si vous auriez fait les choses différemment.